We use a number of analytical tools to perform metabolomic and lipidomic profiling. One of the powerful tools is the High-Performance Chemical Isotope Labeling (CIL) LC-MS Platform (the software for processing CIL LC-MS data is available from www.novamt.com). This platform is very sensitive and provides high metabolome coverage (thousands of metabolites). It is quantitative with high precision and accuracy, as isotope reagents are used for comparative analysis that overcome the matrix and ion suppression effects often associated with conventional LC-MS.
CIL LC-MS Data Analysis Workflow
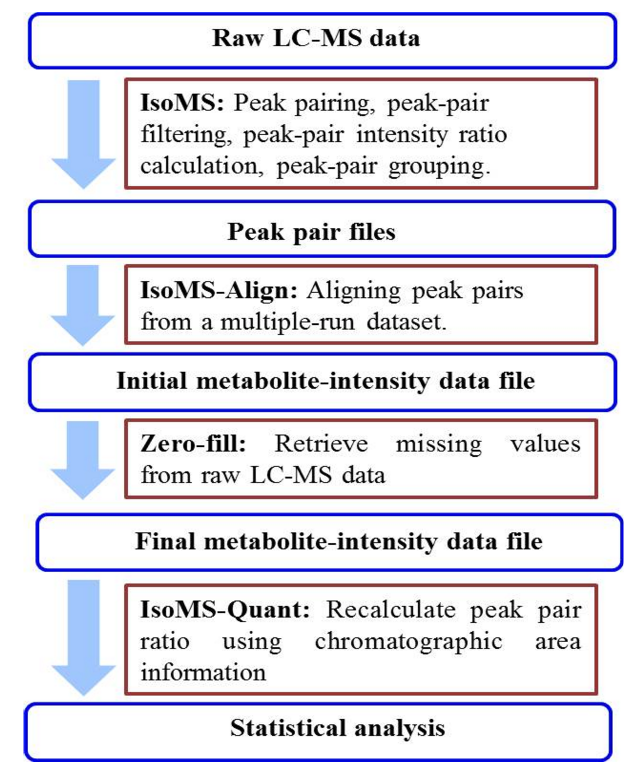
The overall workflow for chemical isotope labeling (CIL) LC-MS metabolomics platform is shown below. Three programs, IsoMS, Zero-fill and IsoMS-Quant, are used in sequence to process the raw LC-MS data in batch mode to generate a metabolite-intensity data file that can be exported into a statistical tool for statistical analysis of the metabolomic profiles.
Metabolite Identification Programs
We have built web-based resource for identification of compounds of interest based on chemical properties of a molecule, such as accurate mass and fragment ion spectral pattern generated by mass spectrometry (MS).
MS Search. This program allows a user to search a query mass to generate a list of possible matches with the metabolites in an evidence-based metabolome library (EML). This library is composed of 8,021 known human endogenous metabolites and their predicted metabolic products (375,809 compounds from one metabolic reaction and 10,583,901 from two reactions). If the MS/MS spectrum of the query mass ion is available, this program also allows the user to interpret the MS/MS spectral pattern against the list of metabolite candidates generated from the mass search to narrow down the list into one or a few unique structures.
PEP Search. This program allows a user to search the MS/MS spectra of both unlabeled and dimethyl labeled peptides to identify and confirm amino acid sequences of di/tripeptides. It also allows a user to search the MS/MS spectrum of an unlabeled peptide to generate a list of possible sequence matches.
M-RT-MS/MS Search. It contains search program to search any one, two or all three parameters (mass, retention time and MS/MS spectrum) against the labeled standards library for possible match.